Clustering using DBSCAN and its extension OPTICS#
The goal here is to see if the clusters that we can see in the UMAP are interesting… And as we will see they are not.
import pandas as pd
import numpy as np
from pathlib import Path
import utils
# dimensionality reduction
from sklearn.decomposition import PCA
from umap import umap_ as UMAP
# graphing
import seaborn as sns
import matplotlib.pyplot as plt
import matplotlib.gridspec as gridspec
import plotly.express as px
import plotly.offline as pyo
pyo.init_notebook_mode()
# clustering
from sklearn.cluster import OPTICS, cluster_optics_dbscan
Preprocessing of the data#
DATA_PATH = Path.cwd() / "../data"
data = {
Path(f).stem: pd.read_csv(f, index_col=0) for f in DATA_PATH.glob("combined_*.csv")
}
print(list(data.keys()))
['combined_metrics_finished_edges', 'combined_metrics_finished_paths', 'combined_metrics_unfinished_edges', 'combined_metrics_unfinished_paths']
features_finished_paths = data["combined_metrics_finished_paths"].reset_index(drop=True)
features_unfinished_paths = data["combined_metrics_unfinished_paths"].reset_index(
drop=True
)
# drop timeout paths
features_unfinished_paths = features_unfinished_paths[features_unfinished_paths["type"] != "timeout"]
features_unfinished_paths
| index | hashedIpAddress | timestamp | durationInSec | path | target | type | backtrack | numberOfPath | path_length | ... | ratio | average_time_on_page | sucessive_pairs | sucessive_pairs_encoded | semantic_similarity | path_degree_abs_sum | path_clustering_abs_sum | path_degree_centrality_abs_sum | path_betweenness_abs_sum | path_closeness_abs_sum | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 4 | 7 | 6d136e371e42474f | 2011-02-07 18:07:50 | 175 | ['4-2-0', 'United_States', 'Agriculture', 'Sug... | Cane_Toad | restart | 0 | 170.0 | 5 | ... | 3 | 35.000000 | [('4-2-0', 'United_States'), ('United_States',... | [0.19640377163887024, 0.2371780127286911, 0.32... | 0.245052 | 7.585732 | -2.590255 | -0.846121 | -2.271594 | -0.927471 |
| 7 | 12 | 781192ee1a708bec | 2011-02-08 03:46:15 | 334 | ['Saint_Kitts_and_Nevis', 'United_Kingdom', 'W... | Sandy_Koufax | restart | 0 | 1.0 | 5 | ... | 3 | 66.800000 | [('Saint_Kitts_and_Nevis', 'United_Kingdom'), ... | [0.48519760370254517, 0.33719098567962646, 0.3... | 0.331910 | 7.578032 | -2.211302 | -0.853821 | -2.267709 | -1.553485 |
| 9 | 14 | 781192ee1a708bec | 2011-02-08 03:55:46 | 403 | ['Symmetry', 'Science', 'Age_of_Enlightenment'... | Scottish_Episcopal_Church | restart | 0 | 3.0 | 6 | ... | 1 | 67.166667 | [('Symmetry', 'Science'), ('Science', 'Age_of_... | [0.33117276430130005, 0.20039768517017365, 0.5... | 0.336678 | 6.702728 | -3.747010 | -1.729125 | -3.174033 | -2.621027 |
| 10 | 16 | 5900aa2d71b99153 | 2011-02-08 04:23:24 | 115 | ['Tasmanian_Devil', 'Dog', 'Postage_stamp', 'W... | Love | restart | 0 | 1.0 | 4 | ... | 3 | 28.750000 | [('Tasmanian_Devil', 'Dog'), ('Dog', 'Postage_... | [0.28771334886550903, 0.27162858843803406, 0.1... | 0.246961 | 4.382027 | -1.092701 | -4.049827 | -6.726992 | -2.428974 |
| 13 | 19 | 2d49dfd2673786bf | 2011-02-08 06:47:04 | 245 | ['Tim_Berners-Lee', 'England', 'Atlantic_Ocean... | Volcanic_pipe | restart | 0 | 2.0 | 6 | ... | 3 | 40.833333 | [('Tim_Berners-Lee', 'England'), ('England', '... | [0.10544373840093613, 0.3065592646598816, 0.41... | 0.339931 | 6.422029 | -3.488079 | -2.009824 | -3.429649 | -2.719443 |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 18974 | 24867 | 109ed71f571d86e9 | 2014-01-15 11:01:43 | 152 | ['Montenegro', 'World_War_II', 'United_States'... | Hurricane_Georges | restart | 0 | 20.0 | 7 | ... | 4 | 21.714286 | [('Montenegro', 'World_War_II'), ('World_War_I... | [0.16399337351322174, 0.3004598617553711, 0.40... | 0.463695 | 6.294228 | -2.251847 | -2.137626 | -3.529575 | -2.658404 |
| 18975 | 24868 | 109ed71f571d86e9 | 2014-01-15 11:36:08 | 72 | ['Wine', 'Georgia_%28country%29', 'Russia'] | History_of_post-Soviet_Russia | restart | 0 | 27.0 | 3 | ... | 2 | 24.000000 | [('Wine', 'Georgia_%28country%29'), ('Georgia_... | [0.12671996653079987, 0.37717318534851074] | 0.251947 | NaN | NaN | NaN | NaN | NaN |
| 18976 | 24869 | 109ed71f571d86e9 | 2014-01-15 12:00:12 | 182 | ['Turks_and_Caicos_Islands', 'United_States', ... | Iraq_War | restart | 0 | 35.0 | 6 | ... | 2 | 30.333333 | [('Turks_and_Caicos_Islands', 'United_States')... | [0.4149491488933563, 0.40031811594963074, 0.39... | 0.421867 | 7.555088 | -1.505405 | -0.876766 | -2.304742 | -1.574654 |
| 18977 | 24870 | 109ed71f571d86e9 | 2014-01-15 12:06:45 | 180 | ['Franz_Kafka', 'Tuberculosis', 'World_Health_... | Cholera | restart | 1 | 37.0 | 6 | ... | 3 | 30.000000 | [('Franz_Kafka', 'Tuberculosis'), ('Tuberculos... | [0.2173137068748474, 0.3479086458683014, 0.420... | 0.368187 | 3.669951 | -3.305998 | -4.761902 | -7.644179 | -4.383847 |
| 18980 | 24874 | 1cf0cbb3281049ab | 2014-01-15 21:54:01 | 352 | ['Mark_Antony', 'Rome', 'Tennis', 'Hawk-Eye', ... | Feather | restart | 0 | 49.0 | 5 | ... | 2 | 70.400000 | [('Mark_Antony', 'Rome'), ('Rome', 'Tennis'), ... | [0.27006441354751587, 0.28349533677101135, 0.2... | 0.237981 | 5.595124 | -1.730546 | -2.836729 | -5.768889 | -2.107167 |
11811 rows × 24 columns
combined_df = pd.concat([features_finished_paths, features_unfinished_paths], axis=0)
combined_df['finished'] = [1] * len(features_finished_paths) + [0] * len(features_unfinished_paths)
combined_df[utils.FEATURES_COLS_USED_FOR_CLUSTERING] = utils.normalize_features(combined_df[utils.FEATURES_COLS_USED_FOR_CLUSTERING])
X = combined_df[utils.FEATURES_COLS_USED_FOR_CLUSTERING].copy().values
#find number of NaNs in each column
nans = np.isnan(X).sum(axis=0)
print(nans)
#drop rows with NaNs
X = X[~np.isnan(X).any(axis=1)]
[0 0 3 0 0 0 0 0 0 0]
Let’s find the dimensionality of the data#
# find the dimensionality of the data
with plt.style.context("dark_background"):
pca = PCA(n_components=X.shape[1])
pca.fit(X)
# plot the explained variance
plt.scatter(range(1, X.shape[1] + 1), pca.explained_variance_ratio_, c="cyan")
plt.xlabel("Number of components", fontsize=16)
plt.ylabel("Explained variance", fontsize=16)
plt.title("Percentage of explained variance\nof each component of the PCA", fontsize=20)
plt.show()
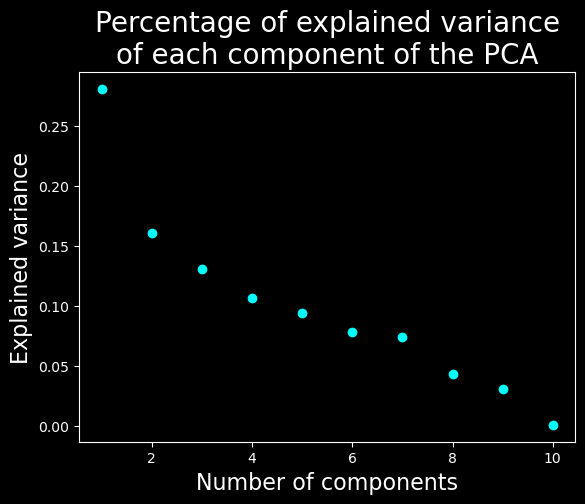
We will keep only the 7 first dimensions
Clustering#
We will run optics on a lower dimension to avoid the curse of the dimensionality#
Let’s find the latent space using a UMAP#
# UMAP
umap = UMAP.UMAP(n_components=7, metric="euclidean")
result_umap_euc = umap.fit_transform(
X
)
Run the clustering in this space using OPTICS or fixed cutoff values with DBSCAN#
#code taken from https://scikit-learn.org/stable/auto_examples/cluster/plot_optics.html
clust = OPTICS(min_samples=50, xi=0.05, min_cluster_size=0.05)
# Run the fit
clust.fit(result_umap_euc)
labels_050 = cluster_optics_dbscan(
reachability=clust.reachability_,
core_distances=clust.core_distances_,
ordering=clust.ordering_,
eps=0.5,
)
labels_200 = cluster_optics_dbscan(
reachability=clust.reachability_,
core_distances=clust.core_distances_,
ordering=clust.ordering_,
eps=2,
)
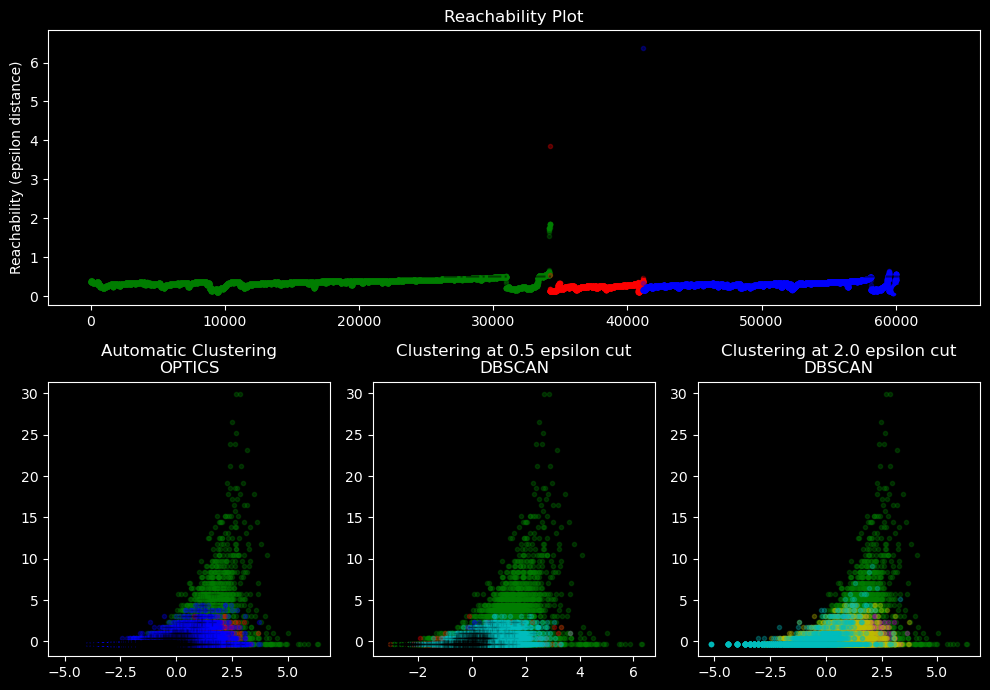
Visualize the results in the plan defined by the 2 first features and draw a reachability plot#
space = np.arange(len(X))
reachability = clust.reachability_[clust.ordering_]
labels = clust.labels_[clust.ordering_]
with plt.style.context("dark_background"):
plt.figure(figsize=(10, 7))
G = gridspec.GridSpec(2, 3)
ax1 = plt.subplot(G[0, :])
ax2 = plt.subplot(G[1, 0])
ax3 = plt.subplot(G[1, 1])
ax4 = plt.subplot(G[1, 2])
# Reachability plot
colors = ["g.", "r.", "b.", "y.", "c."]
for klass, color in zip(range(0, 5), colors):
Xk = space[labels == klass]
Rk = reachability[labels == klass]
ax1.plot(Xk, Rk, color, alpha=0.3)
ax1.plot(space[labels == -1], reachability[labels == -1], "k.", alpha=0.3)
ax1.plot(space, np.full_like(space, 2.0, dtype=float), "k-", alpha=0.5)
ax1.plot(space, np.full_like(space, 0.5, dtype=float), "k-.", alpha=0.5)
ax1.set_ylabel("Reachability (epsilon distance)")
ax1.set_title("Reachability Plot")
# OPTICS
colors = ["g.", "r.", "b.", "y.", "c."]
for klass, color in zip(range(0, 5), colors):
Xk = X[clust.labels_ == klass]
ax2.plot(Xk[:, 0], Xk[:, 1], color, alpha=0.3)
ax2.plot(X[clust.labels_ == -1, 0], X[clust.labels_ == -1, 1], "k+", alpha=0.1)
ax2.set_title("Automatic Clustering\nOPTICS")
# DBSCAN at 0.5
colors = ["g.", "r.", "b.", "c."]
for klass, color in zip(range(0, 4), colors):
Xk = X[labels_050 == klass]
ax3.plot(Xk[:, 0], Xk[:, 1], color, alpha=0.3)
ax3.plot(X[labels_050 == -1, 0], X[labels_050 == -1, 1], "k+", alpha=0.1)
ax3.set_title("Clustering at 0.5 epsilon cut\nDBSCAN")
# DBSCAN at 2.
colors = ["g.", "m.", "y.", "c."]
for klass, color in zip(range(0, 4), colors):
Xk = X[labels_200 == klass]
ax4.plot(Xk[:, 0], Xk[:, 1], color, alpha=0.3)
ax4.plot(X[labels_200 == -1, 0], X[labels_200 == -1, 1], "k+", alpha=0.1)
ax4.set_title("Clustering at 2.0 epsilon cut\nDBSCAN")
plt.tight_layout()
plt.show()
Create a UMAP with 3 dimensions for visualization purposes#
from umap import umap_ as UMAP
# UMAP
umap = UMAP.UMAP(n_components=3, metric="euclidean")
result_umap_euc = umap.fit_transform(
X
)
fig = px.scatter_3d(
result_umap_euc,
x=0,
y=1,
z=2,
color=clust.labels_.astype(str),
title="UMAP, showing OPTICS clustering",
# reduce size points
size_max=0.1,
category_orders={"color": [str(i) for i in range(0, len(np.unique(clust.labels_)))]},
)
fig.update_layout({"plot_bgcolor": "#14181e", "paper_bgcolor": "#14181e"})
fig.update_layout(font_color="white")
fig.update_layout(scene=dict(xaxis=dict(showticklabels=False), yaxis=dict(showticklabels=False), zaxis=dict(showticklabels=False)))
fig.update_layout(legend_title_text="Cluster")
fig.update_layout(legend = dict(bgcolor = 'rgba(0,0,0,0)'))
fig.update_layout(scene=dict(xaxis_title="UMAP 1", yaxis_title="UMAP 2", zaxis_title="UMAP 3"))
fig.update_layout(scene = dict(
xaxis = dict(
backgroundcolor="rgba(0, 0, 0,0)",
# gridcolor="rgba(0, 0, 0,0)", # gridcolor is for logo
showbackground=True,
zerolinecolor="white",),
yaxis = dict(
backgroundcolor="rgba(0, 0, 0,0)",
# gridcolor="rgba(0, 0, 0,0)",
showbackground=True,
zerolinecolor="white"),
zaxis = dict(
backgroundcolor="rgba(0, 0, 0,0)",
# gridcolor="rgba(0, 0, 0,0)",
showbackground=True,
zerolinecolor="white",),),
)
display(fig)
By providing a UMAP in 7 dimension to OPTICS we roughly find the communities that we can see by eye in the 3D UMAP
Visualization of the results#
#plot histogram features colored by cluster
with plt.style.context("dark_background"):
plot_data = combined_df[utils.FEATURES_COLS_USED_FOR_CLUSTERING].copy().dropna()
plot_data["cluster"] = clust.labels_
plot_data["cluster"] = plot_data["cluster"].astype("category")
n_features = len(plot_data.columns) - 1
n_cols = 3
n_rows = int(np.ceil(n_features / n_cols))
fig, axs = plt.subplots(n_rows, n_cols, figsize=(20, 20))
axs = axs.flatten()
for i, col in enumerate(plot_data.columns):
if col == "cluster":
continue
sns.histplot(
data=plot_data,
x=col,
hue="cluster",
ax=axs[i],
stat="density",
common_norm=False,
palette="tab10",
)
utils.set_axis_style(axs, i)
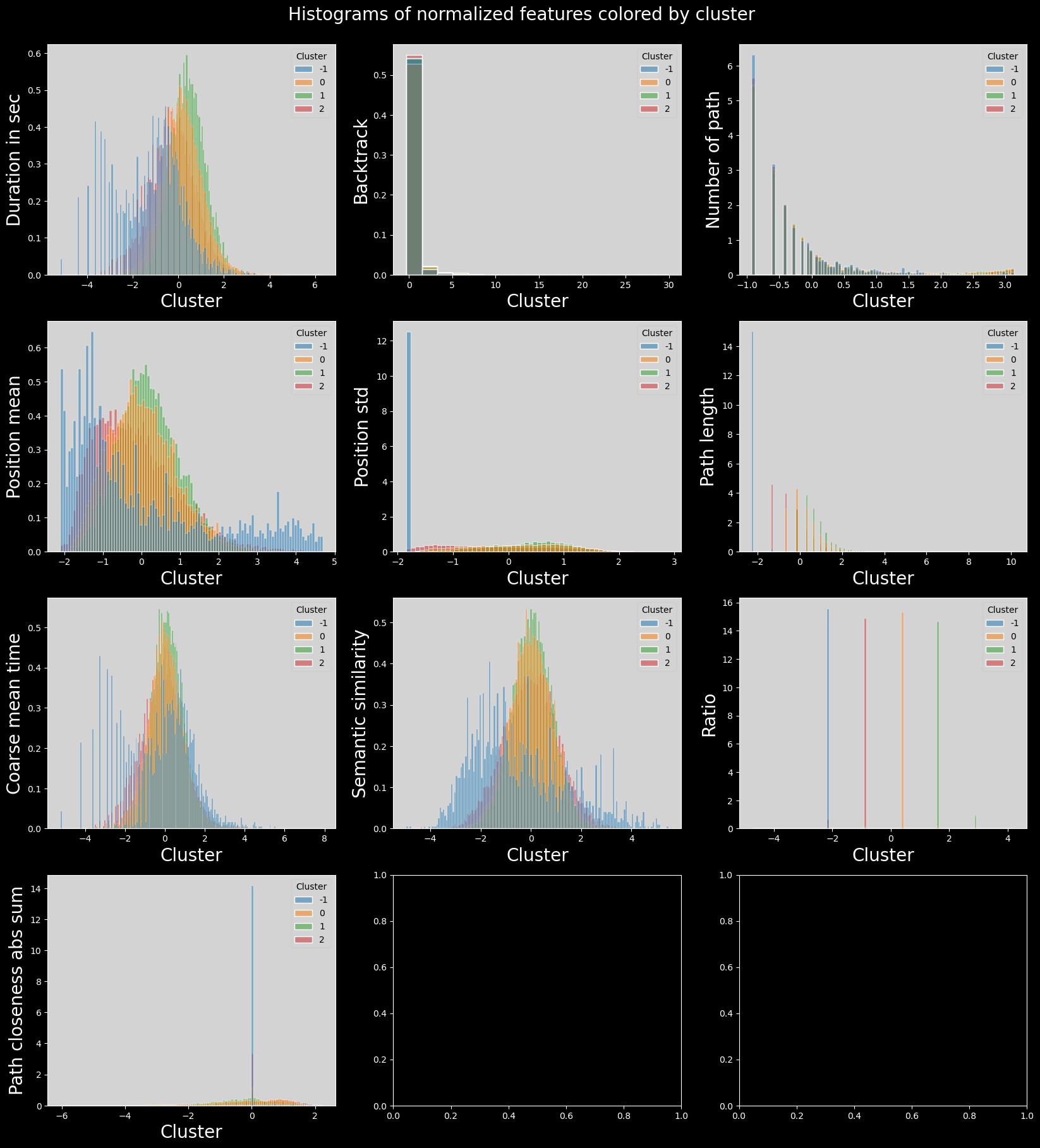
plt.suptitle("Histograms of normalized features colored by cluster", fontsize=20)
plt.subplots_adjust(top=0.95)
plt.show()
with plt.style.context("dark_background"):
combined_df = combined_df[utils.FEATURES_COLS_USED_FOR_CLUSTERING+["finished"]].dropna()
combined_df["finished_normalized"] = (
combined_df["finished"] - combined_df["finished"].mean()
) / combined_df["finished"].std()
plot_data = combined_df[utils.FEATURES_COLS_USED_FOR_CLUSTERING + ["finished_normalized"]].copy()
plot_data["cluster"] = clust.labels_
plot_data["cluster"] = plot_data["cluster"].astype("category")
n_features = len(plot_data.columns) - 1
n_cols = 4
n_rows = int(np.ceil(n_features / n_cols))
fig, axs = plt.subplots(n_rows, n_cols, figsize=(20, 20))
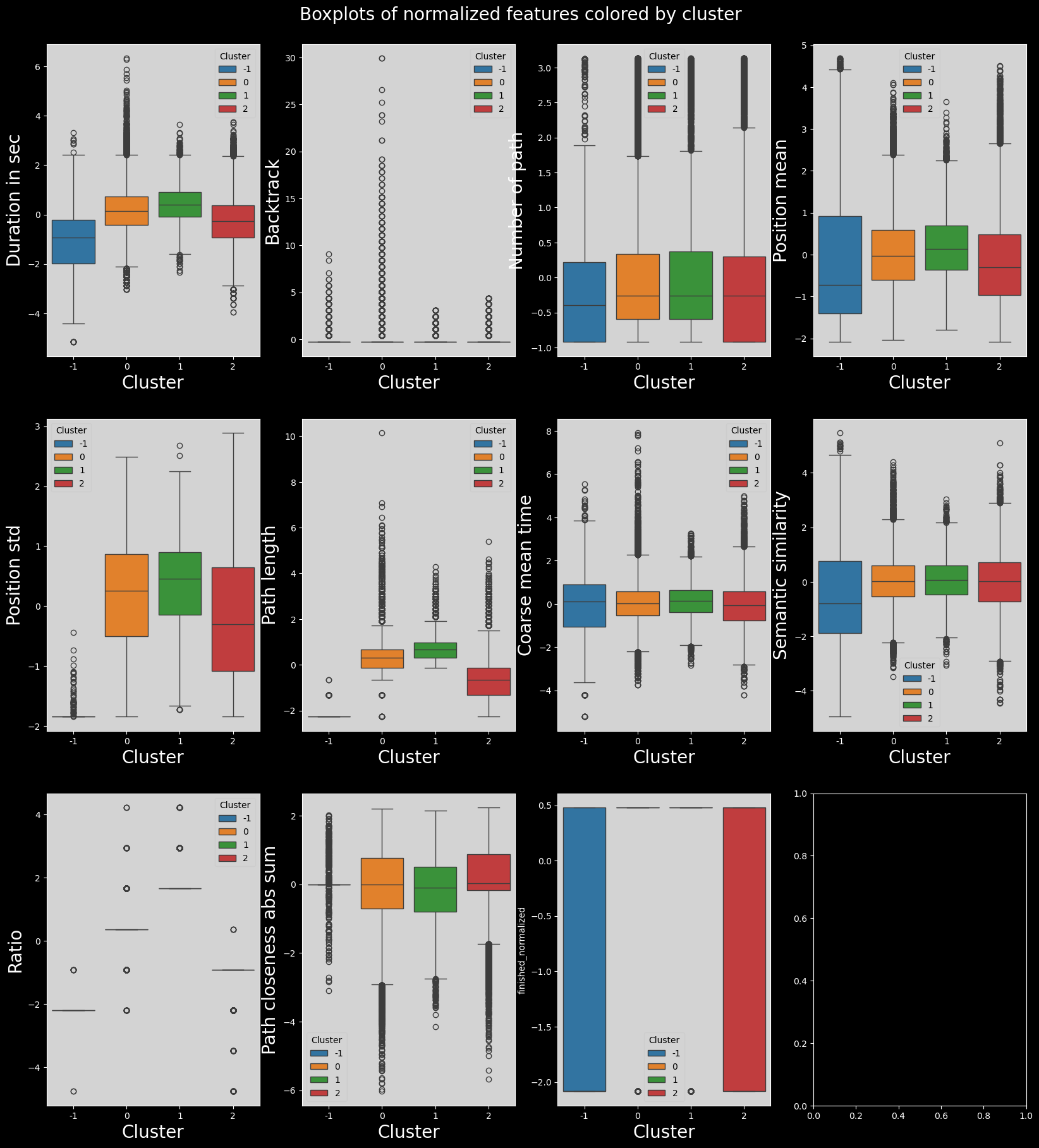
fig.suptitle("Boxplots of normalized features colored by cluster", fontsize=20)
plt.subplots_adjust(top=0.95)
axs = axs.flatten()
for i, col in enumerate(plot_data.columns):
if col == "cluster":
continue
sns.boxplot(data=plot_data, x="cluster", y=col, ax=axs[i], hue="cluster", palette="tab10")
utils.set_axis_style(axs, i)
plt.show()
# heatmap of features means per cluster
feat_labels = utils.get_feature_names_labels()
with plt.style.context("dark_background"):
means = plot_data.groupby(plot_data["cluster"]).mean()
sns.heatmap(means, cmap="coolwarm", annot=True, fmt=".1f", vmin=-1, vmax=1)
plt.title("Heatmap of normalized features\nmeans per cluster", fontsize=25)
plt.ylabel("Cluster", fontsize=20)
plt.xlabel("Feature", fontsize=20)
plt.xticks(labels=feat_labels+["Finished (norm)"], ticks=np.arange(len(feat_labels)+1)+0.5, rotation=90, fontsize=15)
plt.show()
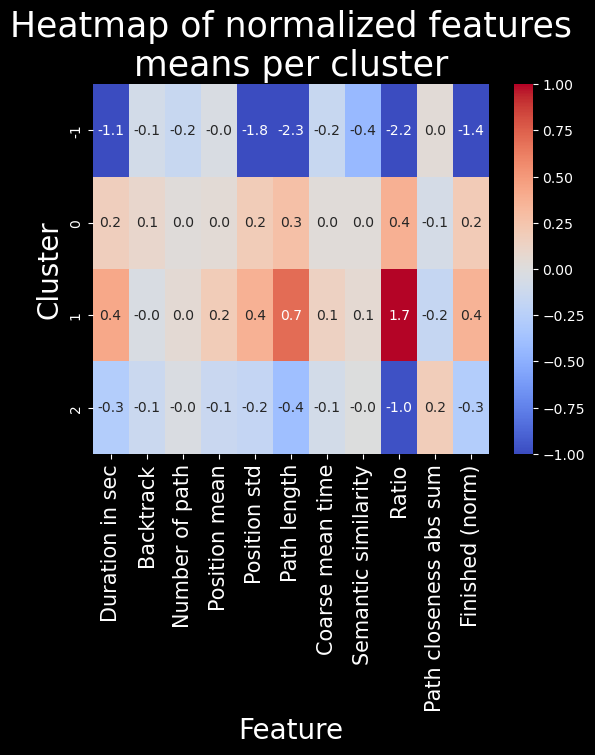
Discussion and conclusion#
Here we can see that the clustering is less interesting. It really makes 4 clusters in which all the features used to do the clustering are either very low, low, high or very high.
